A method based on ontologies to support the automatic management of knowledge about Covid-19
Keywords:
Ontology, knowledge representation, Covid-19, automatic reasoning, decision makingAbstract
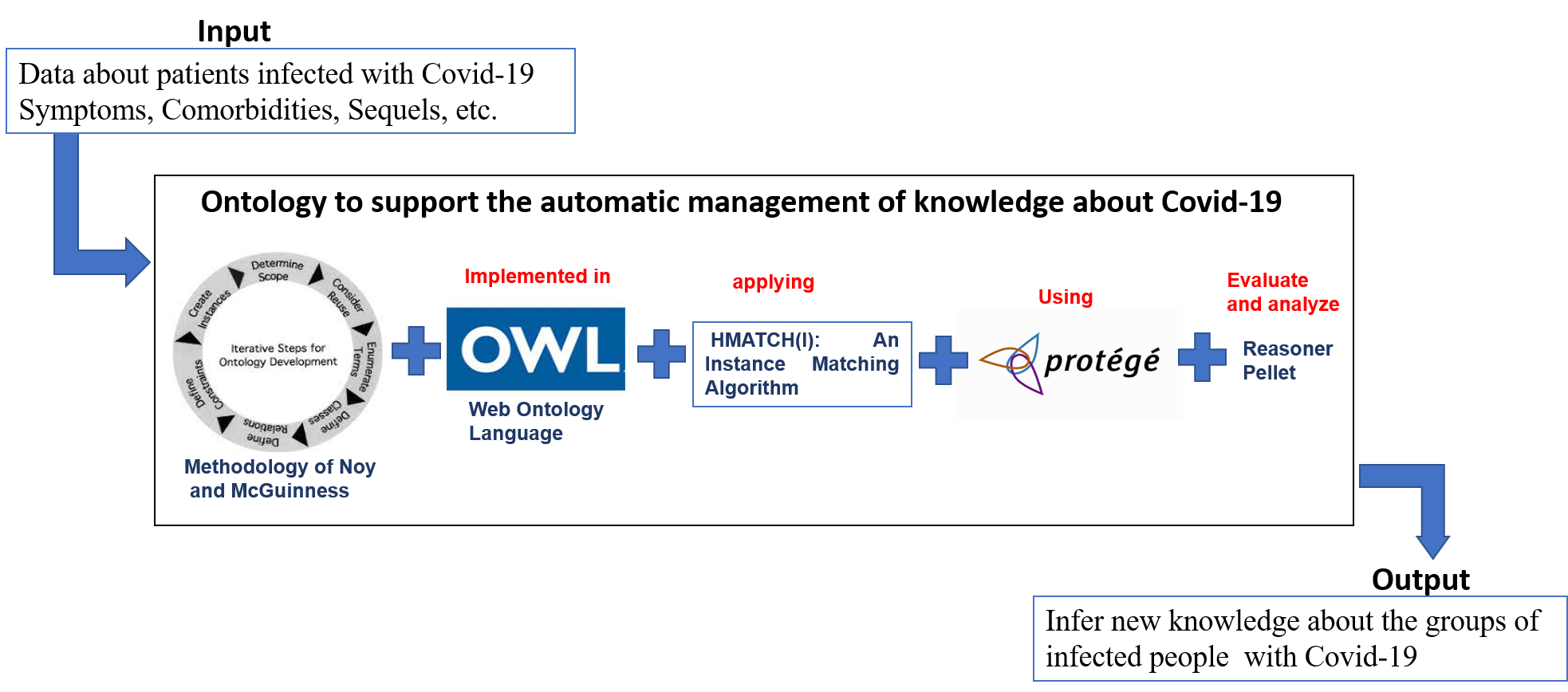
The importance of integration and retrieval of data is growing more and more in the context of post-COVID-19 analysis because of the increased generation of data in research and resources for studying COVID-19. In this context, the analysis of contagion groups that share some specific feature (for example, people that work together, people with the same clinical manifestation, etc.) may offer remarkable insights about COVID-19. For instance, it can be helpful to recognize the behavior of this disease in accordance with the unique characteristics of a group of people. In this regard, ontologies are a widely accepted alternative for representing and analyzing knowledge. Given their benefits, this paper introduces an ontology-based method to formally describe and analyze information about specific groups of people with COVID-19. Since the ontology was specified in OWL, a formal language based on description logics, it enables the consistency of the represented information to be verified and use automatic reasoning to deduce new knowledge. This approach allows modeling a wide variety of characteristics (symptoms, comorbidities, treatments, etc.) of the individuals and consequently use it to infer the general characteristics of the groups that they belong to. Hence, a reasoner can be applied to perform advanced analysis either to identify patterns in these groups or to find similarity with other groups. To demonstrate the applicability of this method, a case study is described. In addition, the ontology was used to represent and analyze the information of a sample of patients extracted from a public dataset. The results demonstrate the capability of the ontology to represent and analyze information about specific groups of people with COVID-19.
Downloads
References
Msemburi, W., Karlinsky, A., Knutson, V., Aleshin, S., Chatterji, S., & Wakefield, J. (2023). The WHO estimates of excess mortality associated with the COVID-19 pandemic. Nature, 613(7942), 130-137. https://doi.org/10.1038/s41586-022-05522-2.
Karlinsky, A., & Kobak, D. (2021). Tracking excess mortality across countries during the COVID-19 pandemic with the World Mortality Dataset. elife, 10, e69336. https://doi.org/10.7554/eLife.69336.
Silega, N., Varén, E., Rogozov, Y., Lapshin, V., & Alekseevich, S., Exploiting an Ontological Model to Study COVID-19 Contagion Chains in Sustainable Smart Cities. Information, 2022. 13(1). https://doi.org/10.3390/info13010040.
Thirumahal, R., Sudha, G., & Shruti, P., Semantic Integration of Heterogeneous Data Sources Using Ontology-Based Domain Knowledge Modeling for Early Detection of COVID-19. SN Computer Science, 2022. 3(6): p. 1-13. https://doi.org/10.1007/s42979-022-01298-4.
Li, G. Improving Biomedical Ontology Matching Using Domain-specific Word Embeddings. In the ACM International Conference Proceeding Series. 2020. https://doi.org/10.1145/3424978.3425102.
Aref, M. M., & Zhou, Z. The Ontology Web Language (OWL) for a multi-agent understanding system. In International Conference on Integration of Knowledge Intensive Multi-Agent Systems, 2005. Waltham, MA, USA, 2005. IEEE, pp. 586-591. https://doi.org/10.1109/KIMAS.2005.1427149.
Suárez, M. C., Gómez, A., Motta, E., & Gangemi, A., Introduction: Ontology Engineering in a Networked World. In: Suárez, M. C., Gómez, A., Motta, E., Gangemi, A. (eds) Ontology Engineering in a Networked World. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-24794-1_1.
Wynants, L., Van Calster, B., Collins, G., Riley, R., Heinze, G., Schuit, E., & van Smeden, M., Prediction models for diagnosis and prognosis of covid-19: systematic review and critical appraisal. The BMJ, 2020. 369(2). https://doi.org/10.1136/bmj.m1328.
Leung, K., Wu, J., & Leung, G., First-wave COVID-19 transmissibility and severity in China outside Hubei after control measures, and second-wave scenario planning: a modelling impact assessment. The Lancet, 2020. 395(10233). https://doi.org/10.1016/S0140-6736(20)30746-7.
Kissler, S., Tedijanto, C., Grad, Y., & Lipsitch, M, Projecting the transmission dynamics of SARS-CoV-2 through the postpandemic period. Science, 2020. 368(6493). https://doi.org/10.1126/science.abb5793.
Bernasconi, A., Canakoglu, A., Pinoli, P., & Ceri, S., Empowering virus sequence research through conceptual modeling. In: Dobbie, G., Frank, U., Kappel, G., Liddle, S.W., Mayr, H.C. (eds) Conceptual Modeling. ER 2020. Lecture Notes in Computer Science, vol 12400. Springer, Cham. https://doi.org/10.1007/978-3-030-62522-1_29.
Ahmed, N., Michelin, R. A., Xue, W., Ruj, S., Kanhere, S. S., & Jha, S., A survey of COVID-19 contact tracing apps. IEEE access, 2020. 8: p. 134577-134601. https://doi.org/10.1109/ACCESS.2020.3010226.
Kaxiras, E. and G. Neofotistos, Multiple epidemic wave model of the COVID-19 pandemic: modeling study. Journal of medical Internet research, 2020. 22(7): p. e20912. https://doi.org/10.2196/20912.
Bastos, S., & Cajueiro, D., Modeling and forecasting the early evolution of the Covid-19 pandemic in Brazil. Scientific Reports, 2020. 10(1): p. 1-10. https://doi.org/10.1038/s41598-020-76257-1.
Sanche, S., Lin, Y., Xu, C., Romero, E., Hengartner, N., & Ke, R., High contagiousness and rapid spread of severe acute respiratory syndrome coronavirus 2. Emerging infectious diseases, 2020. 26(7): p. 1470. https://doi.org/10.3201/eid2607.200282.
Xiang, L., Ma, S., Yu, L., Wang, W., & Yin, Z, Modeling the Global Dynamic Contagion of COVID-19. Frontiers in public health, 2021. 9. https://doi.org/10.3389/fpubh.2021.809987.
Samee, N., El-Kenawy, E., Atteia, G., Jamjoom, M., Ibrahim, A., Abdelhamid, A., & Shams, M., Metaheuristic Optimization Through Deep Learning Classification of COVID-19 in Chest X-Ray Images. Computers, Materials & Continua, 2022. 73(2): 4193–4210. https://doi.org/10.32604/cmc.2022.031147.
Samee, N., Alhussan, A., Ghoneim, V., Atteia, G., Alkanhel, & Kadah, Y., A Hybrid Deep Transfer Learning of CNN-Based LR-PCA for Breast Lesion Diagnosis via Medical Breast Mammograms. Sensors, 2022. 22(13): p. 4938. https://doi.org/10.3390/s22134938.
Nawaz, A., Abbas, Y., Ahmad, T., Mahmoud, N., Rizwan, A., & Samee, N., A Healthcare Paradigm for Deriving Knowledge Using Online Consumers’ Feedback. Multidisc. Digital Publishing Institute. https://doi.org/10.3390/healthcare10081592.
Alhussan, A., Samee, N., Ghoneim, V., & Kadah, Y., Evaluating Deep and Statistical Machine Learning Models in the Classification of Breast Cancer from Digital Mammograms. International Journal of Advanced Computer Science and Applications (IJACSA), 2021. 12(10). https://doi.org/10.14569/IJACSA.2021.0121033.
Atteia, G., Abdel, N., & Zohair, H., DFTSA-Net: Deep Feature Transfer-Based Stacked Autoencoder Network for DME Diagnosis. Entropy, 2021. 23(10): p. 1251. https://doi.org/10.3390/e23101251.
Ahmad, A., Bandara, M., Fahmideh, M., Proper, H., & Soar, J., An Overview of Ontologies and Tool Support for COVID-19 Analytics. Math. Biosci., 328, 108441. https://doi.org/10.1109/EDOCW52865.2021.00026.
Garba, S., Lubuma, J., & Tsanu, B., Modeling the transmission dynamics of the COVID-19 Pandemic in South Africa. Mathematical biosciences, 2020. 328: p. 108441. https://doi.org/10.1016/j.mbs.2020.108441.
Singhal, A., Singh, P., & Joshi, S., Modeling and prediction of COVID-19 pandemic using Gaussian mixture model. Chaos, solitons & fractals, 2020. 138: p. 110023. https://doi.org/10.1016/j.chaos.2020.110023.
Aviv, E., & Aharoni, A, Generalized logistic growth modeling of the Covid-19 pandemic in Asia. Infectious Disease Modelling, 2020. 5: p. 502-509. https://doi.org/10.1016/j.idm.2020.07.003.
Zuo, F., Wang, J., Gao, J., Ozbay, K., Ban, X., Shen, Y., & Iyer, S, An interactive data visualization and analytics tool to evaluate mobility and sociability trends during covid-19. preprint arXiv:2006.14882, 2020. https://doi.org/10.48550/arXiv.2006.14882.
Mahalle, P., Sable, N., & Shinde, G., Data analytics: Covid-19 prediction using multimodal data in Intelligent systems and methods to combat Covid-19. 2020, Springer. https://doi.org/10.1007/978-981-15-6572-4_1.
Wang, C., Ng, C., & Brook, R., Response to COVID-19 in Taiwan: big data analytics, new technology, and proactive testing. Jama, 2020. 323(14): p. 1341-1342. https://doi.org/10.1001/jama.2020.3151.
Dutta, B. & M. DeBellis. Codo: An ontology for collection and analysis of Covid-19 data. in 12th International Joint Conference on Knowledge Discovery, Knowledge Engineering and Knowledge Management. 2020. https://doi.org/10.5220/0010112500760085.
Babcock, S., Beverley, J., Cowell, L., & Smith, B, The Infectious Disease Ontology in the age of COVID-19. Journal of Biomedical Semantics, 2021. 12(1). https://doi.org/10.1186/s13326-021-00245-1.
Sargsyan, A., et al., The COVID-19 ontology. Bioinformatics, 2020. 36(24): p. 5703-5705. https://doi.org/10.1093/bioinformatics/btaa1057.
Cowell, L. & Smith, B., Infectious disease ontology, in Infectious Disease Inform.. 2010, Springer. https://doi.org/10.1007/978-1-4419-1327-2_19.
He, Y., Yu, H., Ong, E., Wang, Y., Liu, Y., & Smith, B, CIDO, a community-based ontology for coronavirus disease knowledge and data integration, sharing, and analysis. Scientific data, 2020. 7(1): p. 181. https://doi.org/10.1038/s41597-020-0523-6.
de Lusignan, S., Liyanage, H., McGagh, D., Jani, B., Bauwens, J., Byford, R., Evans, D., Fahey, T., Greenhalgh, T., Jones, N., Mair, F., Okusi, C., Parimalanathan, V., Pell, J., Sherlock, J., Tamburis, O., Tripathy, M., Ferreira, F., Williams, J., Hobbs, F., Covid-19 surveillance in a primary care sentinel network: In-Pandemic development of an application ontology. JMIR Public Health and Surveillance, 2020. 6(4): p. e21434. https://doi.org/10.2196/21434.
Liu, Y., Chan, W., Wang, Z., Hur, J., Xie, J., Yu, H., & He, Y., Ontological and bioinformatic analysis of anti-coronavirus drugs and their implication for drug repurposing against Covid-19. 2020. https://doi.org/10.20944/preprints202003.0413.v1.
Sayers, S., Li, L., Ong, E., Deng, S., Fu, G., & He, Y., Victors: a web-based knowledge base of virulence factors in human and animal pathogens. Nucleic acids research, 2019. 47(D1): p. D693-D700. https://doi.org/10.1093/nar/gky999.
Sharma, S. and S. Jain, CovidO: an ontology for COVID-19 metadata. The Journal of Supercomputing, 2024. 80(1): p. 1238-1267. https://doi.org/10.1007/s11227-023-05509-4.
Wegner, P., Jose, G., Lage, V., Golriz, S., & Zhang, B., Common data model for COVID-19 datasets. Bioinformatics, 2022. 38(24): p. 5466-5468. https://doi.org/10.1093/bioinformatics/btac651.
Qundus, J.A., Schäfermeier, R., Karam, N., Peikert, S., & Paschke, A., Roc: an ontology for country responses towards covid-19. arXiv:2104.07345, 2021. https://doi.org/10.48550/arXiv.2104.07345.
Narayanasamy, S., Srinivasan, K., Hu, Y., & Huang, K., A Contemporary Review on Utilizing Semantic Web Technologies in Healthcare, Virtual Communities, and Ontology-Based Information Processing Systems. Electronics.2022; 11(3). https://doi.org/10.3390/electronics11030453.
Yang, C., Ambayo, H., De Baets, B., Kolsteren, P., Thanintorn, & Lachat, C, An Ontology to Standardize Research Output of Nutritional Epidemiology: From Paper-Based Standards to Linked Content. Nutrients, 2019. 11(6): p. 1300. https://doi.org/10.3390/nu11061300.
Magumba, M. & P. Nabende. An ontology for generalized disease incidence detection on twitter. In: Martínez de Pisón, F., Urraca, R., Quintián, H., Corchado, E. (eds) Hybrid Artificial Intelligent Systems. HAIS 2017. Lecture Notes in Computer Science, vol 10334. Springer, Cham. https://doi.org/10.1007/978-3-319-59650-1_4.
Amith, M., Fujimoto, K., Mauldin, R., & Tao, C., Friend of a Friend with Benefits ontology (FOAF+): extending a social network ontology for public health. BMC Medical Informatics and Decision Making, 2020. 20(10): p. 1-14. https://doi.org/10.1186/s12911-020-01287-8.
Slimani, T., Ontology development: A comparing study on tools, languages and formalisms. Ind. Jour. of Sci. & Tech., 2015. 8(24): p. 1-12. https://doi.org/10.17485/ijst/2015/v8i34/54249.
Corcho, O., Fernández, M, & Gómez, A., Methodologies, tools and languages for building ontologies. Where is their meeting point? Data & knowledge engineering, 2003. 46(1). https://doi.org/10.1016/S0169-023X(02)00195-7.
Musen, M.A., The protégé project: A look back and a look forward. AI Matters, 2015. 1(4): p. 4-12. https://doi.org/10.1145/2757001.2757003.
Sattar, A., Surin, E., Ahmad, M, & Mahmood, A., Comparative analysis of methodologies for domain ontology development: A systematic review. International Journal of Advanced Computer Science and Applications, 2020. 11(5). https://doi.org/10.14569/IJACSA.2020.0110515.
Castano, S., Ferrara, A., Montanelli, S., & Lorusso, D. Instance Matching for Ontology Population. In Italian Symposium on Advanced Database Systems. 2008. http://hdl.handle.net/2434/233009.
Whetzel, P., Noy, N., Shah, N., Alexander, P., Nyulas, C., Tudorache, T., Musen, M., BioPortal: enhanced functionality via new Web services from the National Center for Biomedical Ontology to access and use ontologies in software applications. Nucleic Acids Res, 2011. 39(suppl_2): p. W541- W545. https://doi.org/10.1093/nar/gkr469.
Poveda, M., A. Gómez, & M. Suárez, Oops! (Ontology Pitfall Scanner!): An on-line tool for ontology evaluation. International Journal on Semantic Web and Information Systems (IJSWIS), 2014. 10(2): p. 7-34. https://doi.org/10.4018/ijswis.2014040102.
Abhilash, C. and K. Mahesh. Ontology-based interestingness in covid-19 data. in Research Conference on Metadata and Semantics Research. 2021. Springer. https://doi.org/10.5220/0010112500760085.