DeepRetinaNet: An Automated AI-Based Framework for Retinal Disease Diagnosis
Keywords:
Retinal image, LSTM, feature fusion, diabetic retinopathy, GlaucomaAbstract
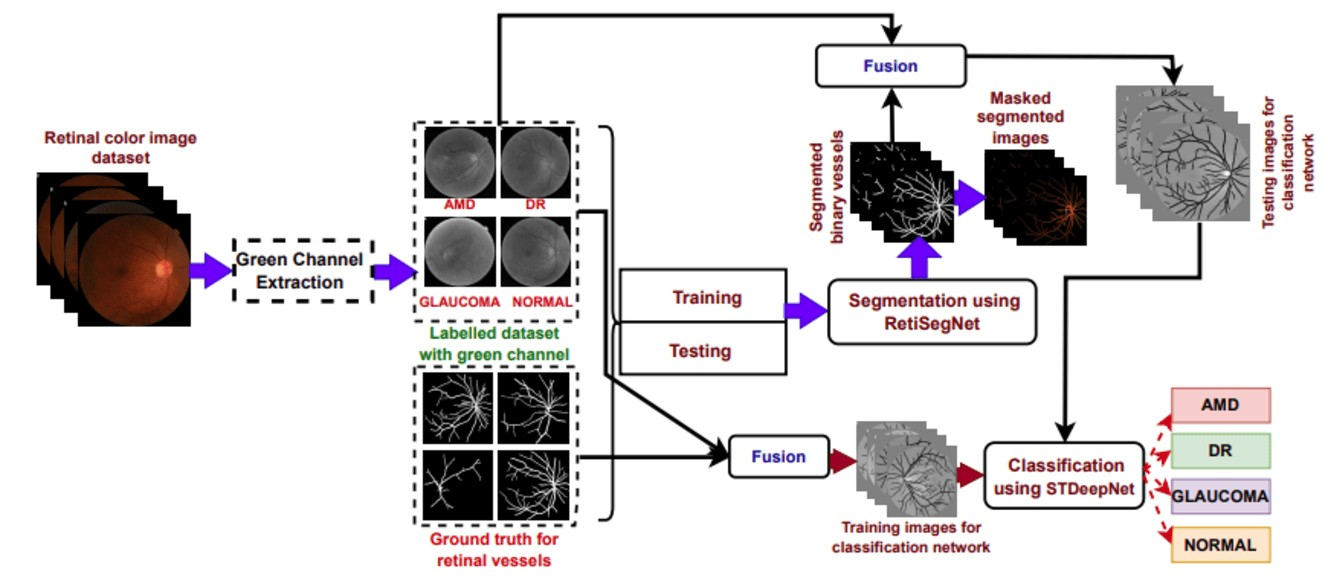
Automated retinal disease diagnosis leveraging cutting-edge computer vision methodologies supports clinicians in the early identification of pathological conditions. This investigation delivers a novel framework, DeepRetinaNet for automating retinal disease diagnosis. The developed DeepRetinaNet model has two stages of novelties, including vessel extraction followed by disease identification. In the vessel extraction stage, the green channel, known for its heightened sensitivity to retinal vascular structures, is extracted from the source images. Subsequently, the vessel extraction network: RetiSegNet, processes these green channel images to extract retinal vessels, generating binary vessel maps. During the fusion phase, the original fundus images are combined with the extracted vessel maps to produce fused representations, encapsulating enriched spatial details from both sources. In the identification stage, these fused images are utilized to train the proposed classification framework: STDeepNet, which incorporates Modified Identity (MI), Modified Convolution (MCONV) blocks, and Long Short-Term Memory (LSTM) layers to effectively identify the diseases. The efficacy of the developed technique is corroborated using visual illustration and objective analysis. Also, the efficiency of the designed framework is verified on six benchmark datasets. The proposed framework demonstrates superior performance compared to 49 state-of-the-art methods, achieving notable accuracy in retinal disease diagnosis.
Downloads
References
M. J. Burton, J. Ramke, A. P. Marques, R. R. Bourne, N. Congdon, I. Jones, B. A. A. Tong, S. Arunga, D. Bachani, C. Bascaran et al., “The lancet global health commission on global eye health: vision beyond
,” The Lancet Global Health, vol. 9, no. 4, pp. 489–551, 2021, doi:10.1016/S2214-109X(20)30488-5.
Z. Qiu, Y. Hu, X. Chen, D. Zeng, Q. Hu, and J. Liu, “Rethinking dual-stream super-resolution semantic learning in medical image segmentation,” IEEE Transactions on Pattern Analysis and Machine Intelligence, vol. 46, no. 1, pp. 451–464, 2024, doi: 10.1109/TPAMI.2023.3322735.
E. ¨Ozbay, “An active deep learning method for diabetic retinopathy detection in segmented fundus images using artificial bee colony algorithm,” Artificial Intelligence Review, vol. 56, no. 4, pp. 3291–3318,
, doi: 10.1007/s10462-022-10231-3.
A. D. Vairamani, “Detection and diagnosis of diseases by feature extraction and analysis on fundus images using deep learning techniques,” in Computational Methods and Deep Learning for Ophthalmology, 2023, doi: 10.1016/B978-0-323-95415-0.00009-7, pp. 211–227.
R. Biswas, A. Vasan, and S. S. Roy, “Dilated deep neural network for segmentation of retinal blood vessels in fundus images,” Iranian Journal of Science and Technology, Transactions of Electrical Engineering, vol. 44, no. 1, pp. 505–518, 2020, doi: 10.1007/s40998-019-00213-7.
A. A. Abdulsahib, M. A. Mahmoud, H. Aris, S. S. Gunasekaran, and M. A. Mohammed, “An automated image segmentation and useful feature extraction algorithm for retinal blood vessels in fundus images,” Electronics, vol. 11, no. 9, p. 1295, 2022, doi: 10.3390/electronics11091295.
Z. Qiu, Y. Hu, H. Li, and J. Liu, “Learnable ophthalmology sam,” arXiv preprint arXiv:2304.13425, 2023.
A. Jayachandran, S. R. Kumar, and T. S. R. Perumal, “Multi-dimensional cascades neural network models for the segmentation of retinal vessels in colour fundus images,” Multimedia Tools and Applications, vol. 82, no. 27, pp. 42 927–42 943, 2023, doi: 10.1007/s11042-023-15133-2.
S. A. David, C. Mahesh, V. D. Kumar, K. Polat, A. Alhudhaif, and M. Nour, “Retinal Blood Vessels and Optic Disc Segmentation Using U-Net,” Mathematical Problems in Engineering, vol. 2022, no. 1, p.
, 2022, doi: 10.1155/2022/8030954.
Y. Xu and Y. Fan, “Dual-channel asymmetric convolutional neural network for an efficient retinal blood vessel segmentation in eye fundus images,” Biocybernetics and Biomedical Engineering, vol. 42, no. 2, pp. 695–706, 2022, doi: 10.1016/j.bbe.2022.05.003.
R. Thanki, “A deep neural network and machine learning approach for retinal fundus image classification,” Healthcare Analytics, vol. 3, p. 100140, 2023, doi: 10.1016/j.health.2023.100140.
I. K. Gupta, A. Choubey, and S. Choubey, “Mayfly optimization with deep learning enabled retinal fundus image classification model,” Computers and Electrical Engineering, vol. 102, p. 108176, 2022, doi:
1016/j.compeleceng.2022.108176.
S. Srinivasan, R. Nagarnaidu Rajaperumal, S. K. Mathivanan, P. Jayagopal, S. Krishnamoorthy, and S. Kardy, “Detection and grade classification of diabetic retinopathy and adult vitelliform macular dystrophy based on ophthalmoscopy images,” Electronics, vol. 12, no. 4, p. 862, 2023, doi: 10.3390/electronics12040862.
N. Sengar, R. C. Joshi, M. K. Dutta, and R. Burget, “EyeDeep-Net: a multi-class diagnosis of retinal diseases using deep neural network,” Neural Computing and Applications, vol. 35, no. 14, pp. 10 551–10 571, 2023, doi: 10.1007/s00521-023-08249-x.
S. Kumar and B. Kumar, “Automatic early glaucoma detection by extracting parapapillary atrophy and optic disc from fundus image using svm,” Multimedia Tools and Applications, vol. 81, no. 10, pp. 13 513–13 535, 2022, doi: 10.1007/s11042-021-11023-7.
F. Li, Y. Wang, T. Xu, L. Dong, L. Yan, M. Jiang, X. Zhang, H. Jiang, Z. Wu, and H. Zou, “Deep learning-based automated detection for diabetic retinopathy and diabetic macular oedema in retinal fundus photographs,” Eye, vol. 36, no. 7, pp. 1433–1441, 2022, doi: 10.1038/s41433-021-01552-8.
Y. Kumar and B. Gupta, “Retinal image blood vessel classification using hybrid deep learning in cataract diseased fundus images,” Biomedical Signal Processing and Control, vol. 84, p. 104776, 2023, doi: 10.1016/j.bspc.2023.104776.
K. Susheel Kumar and N. Pratap Singh, “Identification of retinal diseases based on retinal blood vessel segmentation using dagum pdf and feature-based machine learning,” The Imaging Science Journal, vol. 71, no. 5,pp. 425–445, 2023, doi: 10.1080/13682199.2023.2183319.
R. C. Joshi, A. K. Sharma, and M. K. Dutta, “VisionDeep-AI: Deep learning-based retinal blood vessels segmentation and multi-class classification framework for eye diagnosis,” Biomedical Signal Processing
and Control, vol. 94, p. 106273, 2024, doi: 10.1016/j.bspc.2024.106273.
D. Nagpal, N. Alsubaie, B. O. Soufiene, M. S. Alqahtani, M. Abbas, and H. M. Almohiy, “Automatic detection of diabetic hypertensive retinopathy in fundus images using transfer learning,” Applied Sciences, vol. 13, no. 8, p. 4695, 2023, doi: 10.3390/app13084695.
K. Jin, X. Huang, J. Zhou, Y. Li, Y. Yan, Y. Sun, Q. Zhang, Y. Wang, and J. Ye, “FIVES: A fundus image dataset for artificial Intelligence based vessel segmentation,” Scientific data, vol. 9, no. 1, p. 475, 2022,
doi:10.1038/s41597-022-01564-3.
M. M. Fraz, P. Remagnino, A. Hoppe, B. Uyyanonvara, A. R. Rudnicka, C. G. Owen, and S. A. Barman, “An ensemble classification-based approach applied to retinal blood vessel segmentation,” IEEE Transactions on Biomedical Engineering, vol. 59, no. 9, pp. 2538–2548, 2012, doi: 10.1109/TBME.2012.2205687.
S. Pachade, P. Porwal, D. Thulkar, M. Kokare, G. Deshmukh, V. Sahasrabuddhe, L. Giancardo, G. Quellec, and F. M´eriaudeau, “Retinal fundus multi-disease image dataset (RFMiD): a dataset for multi-
disease detection research,” Data, vol. 6, no. 2, p. 14, 2021, doi: 10.3390/data6020014.
D. S. W. Ting, L. R. Pasquale, L. Peng, J. P. Campbell, A. Y. Lee, R. Raman, G. S. W. Tan, L. Schmetterer, P. A. Keane, and T. Y. Wong, “Artificial intelligence and deep learning in ophthalmology,” British
Journal of Ophthalmology, vol. 103, no. 2, pp. 167–175, 2019, doi: 10.1136/bjophthalmol-2018-313173.
H. Xiong, F. Long, M. S. Alam, and J. Sang, “Multi-GlaucNet: A multi-task model for optic disc segmentation, blood vessel segmentation and glaucoma detection,” Biomedical Signal Processing and Control, vol. 99, p. 106850, 2025, doi: 10.1016/j.bspc.2024.106850.
S. Al-Fahdawi, A. S. Al-Waisy, D. Q. Zeebaree, R. Qahwaji, H. Natiq, M. A. Mohammed, J. Nedoma, R. Martinek, and M. Deveci, “Fundus-DeepNet: Multi-label deep learning classification system for
enhanced detection of multiple ocular diseases through data fusion of fundus images,” Information Fusion, vol. 102, p. 102059, 2024, doi: 10.1016/j.inffus.2023.102059.
J. Lin, X. Huang, H. Zhou, Y. Wang, and Q. Zhang, “Stimulus-guided adaptive transformer network for retinal blood vessel segmentation in fundus images,” Medical Image Analysis, vol. 89, p. 102929, 2023, doi: 10.1016/j.media.2023.102929.
W. Zhou, W. Bai, J. Ji, Y. Yi, N. Zhang, and W. Cui, “Dual-path multi-scale context dense aggregation network for retinal vessel segmentation,” Computers in Biology and Medicine, vol. 164, p. 107269, 2023, doi: 10.1016/j.compbiomed.2023.107269.
Y. Li, Y. Zhang, J.-Y. Liu, K. Wang, K. Zhang, G.-S. Zhang, X.-F. Liao, and G. Yang, “Global transformer and dual local attention network via deep-shallow hierarchical feature fusion for retinal vessel segmentation,” IEEE Transactions on Cybernetics, vol. 53, no. 9, pp. 5826–5839, 2022, doi: 10.1109/TCYB.2022.3194099.
Y. Yeganeh, A. Farshad, G. Guevercin, A. Abu-zer, R. Xiao, Y. Tang, E. Adeli, and N. Navab, “Scope: Structural continuity preservation for medical image segmentation,” arXiv preprint arXiv:2304.14572, 2023, doi: 10.48550/arXiv.2304.14572.
H. Zhang, W. Ni, Y. Luo, Y. Feng, R. Song, and X. Wang, “TUnet-LBF: Retinal fundus image fine segmentation model based on transformer Unet network and LBF,” Computers in Biology and Medicine, vol. 159, p. 106937, 2023, doi: 10.1016/j.compbiomed.2023.106937.
J. Ryu, M. U. Rehman, I. F. Nizami, and K. T. Chong, “SegR-Net: A deep learning framework with multi-scale feature fusion for robust retinal vessel segmentation,” Computers in Biology and Medicine, vol. 163, p. 107132, 2023, doi: 10.1016/j.compbiomed.2023.107132.
Y. Xie, J. Shang, Q. Yang, X. Qian, H. Zhang, and X. Tang, “ARSA- UNet: Atrous residual network based on structure-adaptive model for retinal vessel segmentation,” Biomedical Signal Processing and Control, vol. 96, p. 106595, 2024, doi: 10.1016/j.bspc.2024.106595.
Y. Zhou, H. Yu, and H. Shi, “Study group learning: Improving retinal vessel segmentation trained with noisy labels,” in Medical Image Computing and Computer Assisted Intervention–MICCAI, 2021, doi:
1007/978-3-030-87193-2 6, pp. 57–67.
M. T. Sadrabadi and H. Agahi, “Comparative analysis of retinal vessel segmentation utilising convolutional and transformer-based architectures,” pp. 1–12, 2024, doi: dx.doi.org/10.2139/ssrn.5003364.
R. Li, M. Li, J. Li, and Y. Zhou, “Connection sensitive attention U-NET for accurate retinal vessel segmentation,” arXiv preprint arXiv:1903.05558, 2019, doi: 10.48550/arXiv.1903.05558.
Z. Fan, J. Mo, B. Qiu, W. Li, G. Zhu, C. Li, J. Hu, Y. Rong, and X. Chen, “Accurate retinal vessel segmentation via octave convolution neural network,” arXiv preprint arXiv:1906.12193, 2019, doi:
48550/arXiv.1906.12193.
S. A. Kamran, K. F. Hossain, A. Tavakkoli, S. L. Zuckerbrod, K. M. Sanders, and S. A. Baker, “RV-GAN: Segmenting retinal vascular structure in fundus photographs using a novel multi-scale generative ad-
versarial network,” in Medical image computing and computer assisted intervention–MICCAI, 2021, doi: 10.1007/978-3-030-87237-3 4.
X. Sun, H. Fang, Y. Yang, D. Zhu, L. Wang, J. Liu, and Y. Xu, “Robust retinal vessel segmentation from a data augmentation perspective,” in Ophthalmic Medical Image Analysis: 8th International Workshop, OMIA 2021, Held in Conjunction with MICCAI, 2021, doi: 10.1007/978-3-030-87000-3 20.
F. T. J. Faria, M. B. Moin, P. Debnath, A. I. Fahim, and F. M. Shah, “Explainable convolutional neural networks for retinal fundus classification and cutting-edge segmentation models for retinal blood
vessels from fundus images,” arXiv preprint arXiv:2405.07338, 2024.
M. Tan and Q. Le, “EfficientNetV2: Smaller models and faster training,” in International conference on machine learning. PMLR, 2021, pp. 10 096–10 106.
N. Park and S. Kim, “How do vision transformers work?” arXiv preprint arXiv:2202.06709, 2022.
M. Tan and Q. Le, “EfficientNet: Rethinking model scaling for convolutional neural networks,” in Proceedings of the 36th International Conference on Machine Learning, ser. Proceedings of Machine Learning Research, K. Chaudhuri and R. Salakhutdinov, Eds., vol. 97. PMLR, 09–15 Jun 2019, pp. 6105–6114.
C. Szegedy, V. Vanhoucke, S. Ioffe, J. Shlens, and Z. Wojna, “Rethinking the inception architecture for computer vision,” in Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2016, pp. 2818–2826.
G. Huang, Z. Liu, L. Van Der Maaten, and K. Q. Weinberger, “Densely connected convolutional networks,” in Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2017, pp. 4700–4708.
K. He, X. Zhang, S. Ren, and J. Sun, “Deep residual learning for image recognition,” in Proceedings of the IEEE conference on Computer Vision and Pattern Recognition, 2016, pp. 770–778.
S. Xie, R. Girshick, P. Doll´ar, Z. Tu, and K. He, “Aggregated residual transformations for deep neural networks,” in Proceedings of the IEEE conference on Computer Vision and Pattern Recognition, 2017, pp. 1492–1500.
X. Xia, Y. Li, G. Xiao, K. Zhan, J. Yan, C. Cai, Y. Fang, and G. Huang, “Benchmarking deep models on retinal fundus disease diagnosis and a large-scale dataset,” Signal Processing: Image Communication, vol. 127, pp. 117–131, 2024, doi: 10.1016/j.image.2024.117151.
C. Priyadharsini et al., “Deep hybrid architecture with stacked ensemble learning for binary classification of retinal disease,” Results in Engineering, vol. 24, pp. 103–122, 2024, doi: 10.1016/j.rineng.2024.103219.
N. Gour and P. Khanna, “Multi-class multi-label ophthalmological disease detection using transfer learning based convolutional neural network,” Biomedical Signal Processing and Control, vol. 66, p. 102329, 2021, doi: 10.1016/j.bspc.2020.102329.
N. Li, T. Li, C. Hu, K. Wang, and H. Kang, “A benchmark of ocular disease intelligent recognition: One shot for multi-disease detection,” in Benchmarking, Measuring, and Optimizing: Third Bench Council
International Symposium, Bench 2021, pp. 177–193.
X. Ou, L. Gao, X. Quan, H. Zhang, J. Yang, and W. Li, “BFENet: A two-stream interaction CNN method for multi-label ophthalmic diseases classification with bilateral fundus images,” Computer Meth-
ods and Programs in Biomedicine, vol. 219, p. 106739, 2022, doi: 10.1016/j.cmpb.2022.106739.
J. He, C. Li, J. Ye, Y. Qiao, and L. Gu, “Multi-label ocular disease classification with a dense correlation deep neural network,” Biomedical Signal Processing and Control, vol. 63, p. 102167, 2021, doi:
1016/j.bspc.2020.102167.