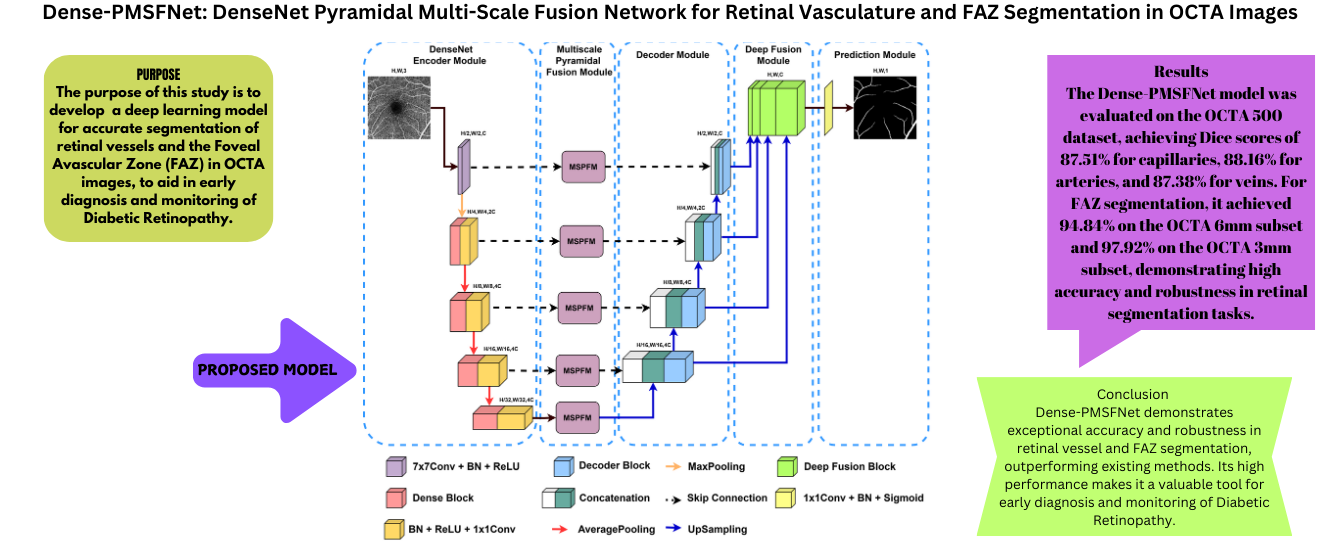
Dense-PMSFNet: DenseNet Pyramidal Multi-Scale Fusion Network for Retinal Vasculature and FAZ Segmentation in OCTA Images
Keywords:
Optical Coherence Tomography Angiography, Artificial Intelligence, vein, artery, capillary, FAZ, densenet encoder, multiscale pyramidal fusion module (MSPFM), deep fusion, deep learning.Abstract
Diabetic retinopathy (DR) is a condition that leads to damage to the retina due to high blood sugar levels in the eyes caused by prolonged diabetes. People of working age in developing countries are particularly at risk of developing diabetic retinopathy, which causes permanent vision loss in diabetic patients. Diagnosis involves maintaining the current level of vision of the patient, as the disease can cause blindness. Optical Coherence Tomography Angiography (OCTA) is an advanced eye imaging technique that offers a detailed view of the structures of retinal blood vessels and effectively treats various eye conditions. Therefore, it is crucial to accurately identify and separate the capillary, artery, vein, and Foveal Avascular Zone (FAZ) from OCTA images. The DenseNet Pyramidal MultiScale Fusion Network (Dense-PMSFNet) is introduced in this article as a new segmentation network based on deep learning. It utilizes the DenseNet Encoder to integrate local semantic contextual information, the Multi-Scale Pyramidal Fusion Module (MSPFM) to effectively merge local features at different scales, and Deep Fusion to enhance the representation of multiscale decoder outputs. Through extensive experimentation on four distinct segmentation tasks involving capillary, artery, vein, and FAZ from OCTA images, it has demonstrated superior performance for the quantitative metrics such as Dice coefficient (DSC), Accuracy (ACC), Intersection Over Union (IoU), Specificity (SP) and Sensitivity (SE) in comparison to state-of-the-art methods.
Downloads
References
M. Z. Atwany, A. H. Sahyoun, and M. Yaqub, “Deep
learning techniques for diabetic retinopathy classifica-
tion: A survey,” IEEE Access, vol. 10, pp. 28 642–
655, 2022. DOI: 10.1109/ACCESS.2022.3157632.
R. F. Spaide, J. G. Fujimoto, N. K. Waheed, S. R.
Sadda, and G. Staurenghi, “Optical coherence tomogra-
phy angiography,” Progress in retinal and eye research,
vol. 64, pp. 1–55, 2018. DOI: https://doi.org/10.1016/j.
preteyeres.2017.11.003.
Z. Sun, D. Yang, Z. Tang, D. S. Ng, and C. Y. Cheung,
“Optical coherence tomography angiography in diabetic
retinopathy: An updated review,” Eye, vol. 35, no. 1,
pp. 149–161, 2021. DOI: https : / / doi . org / 10 . 1038 /
s41433-020-01233-y.
M. M. Abdelsalam, “Effective blood vessels reconstruc-
tion methodology for early detection and classification
of diabetic retinopathy using octa images by artificial
neural network,” Informatics in Medicine Unlocked,
vol. 20, p. 100 390, 2020. DOI: https : / / doi . org / 10 .
/j.imu.2020.100390.
M. K. Ikram, Y. T. Ong, C. Y. Cheung, and T. Y. Wong,
“Retinal vascular caliber measurements: Clinical sig-
nificance, current knowledge and future perspectives,”
Ophthalmologica, vol. 229, no. 3, pp. 125–136, 2013.
DOI: https://doi.org/10.1159/000342158.
C. Y. Cheung, M. K. Ikram, R. Klein, and T. Y. Wong,
“The clinical implications of recent studies on the
structure and function of the retinal microvasculature
in diabetes,” Diabetologia, vol. 58, pp. 871–885, 2015.
DOI: https://doi.org/10.1007/s00125-015-3511-1.
C. Y.-l. Cheung, C. Sabanayagam, A. K.-p. Law, et
al., “Retinal vascular geometry and 6 year incidence
and progression of diabetic retinopathy,” Diabetologia,
vol. 60, pp. 1770–1781, 2017. DOI: https://doi.org/10.
/s00125-017-4333-0.
Y. Jia, S. T. Bailey, T. S. Hwang, et al., “Quantitative
optical coherence tomography angiography of vascular
abnormalities in the living human eye,” Proceedings of
the National Academy of Sciences, vol. 112, no. 18,
E2395–E2402, 2015. DOI: https://doi.org/10.1073/pnas.
M. Guo, M. Zhao, A. M. Cheong, et al., “Can deep
learning improve the automatic segmentation of deep
foveal avascular zone in optical coherence tomography
angiography?” Biomedical Signal Processing and Con-
trol, vol. 66, p. 102 456, 2021. DOI: https://doi.org/10.
/j.bspc.2021.102456.
Y. Liu, L. Zuo, Y. He, et al., “Octa segmentation with
limited training data using disentangled representation
learning,” in Deep Learning for Medical Image Analy-
sis, Elsevier, 2024, pp. 451–469. DOI: https://doi.org/
1016/B978-0-32-385124-4.00027-1.
Y. Xu, X. Xu, L. Jin, et al., “Partially-supervised
learning for vessel segmentation in ocular images,”
in Medical Image Computing and Computer Assisted
Intervention–MICCAI 2021, Springer, 2021, pp. 271–
DOI: https://doi.org/10.1007/978- 3- 030- 87193-
26.
D. Lin, J. Dai, J. Jia, K. He, and J. Sun, “Scrib-
blesup: Scribble-supervised convolutional networks for
semantic segmentation,” in Proceedings of the IEEE
conference on computer vision and pattern recognition,
, pp. 3159–3167. DOI: 10.1109/CVPR.2016.344.
A. Chinkamol, V. Kanjaras, P. Sawangjai, et al., “Oc-
tave: 2d en face optical coherence tomography angiog-
raphy vessel segmentation in weakly-supervised learn-
ing with locality augmentation,” IEEE Transactions on
Biomedical Engineering, 2022. DOI: 10.1109/TBME.
3232102.
M. Li, K. Huang, Q. Xu, et al., “Octa-500: A retinal
dataset for optical coherence tomography angiography
study,” Medical Image Analysis, vol. 93, p. 103 092,
DOI: https : / / doi . org / 10 . 1016 / j . media . 2024 .
R. Mirshahi, P. Anvari, H. Riazi-Esfahani, M. Sar-
darinia, M. Naseripour, and K. G. Falavarjani, “Foveal
avascular zone segmentation in optical coherence to-
mography angiography images using a deep learning
approach,” Scientific reports, vol. 11, no. 1, p. 1031,
DOI: https://doi.org/10.1038/s41598-020-80058-
x.
T. T. Hormel, T. S. Hwang, S. T. Bailey, D. J. Wilson,
D. Huang, and Y. Jia, “Artificial intelligence in oct
angiography,” Progress in Retinal and Eye Research,
vol. 85, p. 100 965, 2021. DOI: https://doi.org/10.1016/
j.preteyeres.2021.100965.
L. Mou, Y. Zhao, H. Fu, et al., “Cs2-net: Deep learning
segmentation of curvilinear structures in medical imag-
ing,” Medical image analysis, vol. 67, p. 101 874, 2021.
DOI: https://doi.org/10.1016/j.media.2020.101874.
O. Ronneberger, P. Fischer, and T. Brox, “U-net: Convo-
lutional networks for biomedical image segmentation,”
in Medical image computing and computer-assisted
intervention–MICCAI 2015, Springer, 2015, pp. 234–
DOI: https://doi.org/10.1007/978- 3- 319- 24574-
28.
Z. Zhou, M. M. R. Siddiquee, N. Tajbakhsh, and
J. Liang, “Unet++: Redesigning skip connections to
exploit multiscale features in image segmentation,”
IEEE transactions on medical imaging, vol. 39, no. 6,
pp. 1856–1867, 2019. DOI: 10 . 1109 / TMI . 2019 .
K. Huang, N. Su, X. Ma, et al., “Choroidal vessel
segmentation in sd-oct with 3d shape-aware adversarial
networks,” Biomedical Signal Processing and Control,
vol. 84, p. 104 982, 2023. DOI: https://doi.org/10.1016/
j.bspc.2023.104982.
B. Dashtbozorg, A. M. Mendonc¸a, and A. Campilho,
“An automatic graph-based approach for artery/vein
classification in retinal images,” IEEE Transactions on
Image Processing, vol. 23, no. 3, pp. 1073–1083, 2013.
DOI: 10.1109/TIP.2013.2263809.
Y. Zhao, J. Xie, H. Zhang, et al., “Retinal vascular
network topology reconstruction and artery/vein classi-
fication via dominant set clustering,” IEEE transactions
on medical imaging, vol. 39, no. 2, pp. 341–356, 2019.
DOI: 10.1109/TMI.2019.2926492.
S. Zhang, R. Zheng, Y. Luo, et al., “Simultaneous
arteriole and venule segmentation of dual-modal fun-
dus images using a multi-task cascade network,” IEEE
Access, vol. 7, pp. 57 561–57 573, 2019. DOI: 10.1109/
ACCESS.2019.2914319.
M. Alam, D. Toslak, J. I. Lim, and X. Yao, “Color
fundus image guided artery-vein differentiation in op-
tical coherence tomography angiography,” Investiga-
tive ophthalmology & visual science, vol. 59, no. 12,
pp. 4953–4962, 2018. DOI: https : / / doi . org / 10 . 1167 /
iovs.18-24831.
M. Alam, D. Le, T. Son, J. I. Lim, and X. Yao, “Av-net:
Deep learning for fully automated artery-vein classifi-
cation in optical coherence tomography angiography,”
Biomedical optics express, vol. 11, no. 9, pp. 5249–
, 2020. DOI: https://doi.org/10.1364/BOE.399514.
J. Xie, Y. Liu, Y. Zheng, et al., “Classification of retinal
vessels into artery-vein in oct angiography guided by
fundus images,” in Medical Image Computing and Com-
puter Assisted Intervention–MICCAI 2020, Springer,
, pp. 117–127. DOI: https://doi.org/10.1007/978-
-030-59725-2 12.
Y. Zheng, J. S. Gandhi, A. N. Stangos, C. Campa, D. M.
Broadbent, and S. P. Harding, “Automated segmentation
of foveal avascular zone in fundus fluorescein angiog-
raphy,” Investigative ophthalmology & visual science,
vol. 51, no. 7, pp. 3653–3659, 2010. DOI: https://doi.
org/10.1167/iovs.09-4935.
M. D´ıaz, J. Novo, P. Cutr´ın, F. G´omez-Ulla, M. G.
Penedo, and M. Ortega, “Automatic segmentation of
the foveal avascular zone in ophthalmological oct-a
images,” PloS one, vol. 14, no. 2, e0212364, 2019. DOI:
https://doi.org/10.1371/journal.pone.0212364.
M. Li, Y. Chen, Z. Ji, et al., “Image projection network:
d to 2d image segmentation in octa images,” IEEE
Transactions on Medical Imaging, vol. 39, no. 11,
pp. 3343–3354, 2020. DOI: 10 . 1109 / TMI . 2020 .
G. Huang, Z. Liu, L. Van Der Maaten, and K. Q. Wein-
berger, “Densely connected convolutional networks,” in
Proceedings of the IEEE conference on computer vision
and pattern recognition, 2017, pp. 4700–4708. DOI: 10.
/CVPR.2017.243.
A. G. Roy, N. Navab, and C. Wachinger, “Recalibrating
fully convolutional networks with spatial and channel
“squeeze and excitation” blocks,” IEEE Transactions on
Medical Imaging, vol. 38, no. 2, pp. 540–549, 2019.
DOI: 10.1109/TMI.2018.2867261.
H. Zhao, J. Shi, X. Qi, X. Wang, and J. Jia, “Pyramid
scene parsing network,” in Proceedings of the IEEE
conference on computer vision and pattern recognition,
, pp. 2881–2890. DOI: 10.1109/CVPR.2017.660.
J. Bertels, T. Eelbode, M. Berman, et al., “Optimizing
the dice score and jaccard index for medical image
segmentation: Theory and practice,” in Medical Im-
age Computing and Computer Assisted Intervention–
MICCAI 2019, Springer, 2019, pp. 92–100. DOI: https:
//doi.org/10.1007/978-3-030-32245-8 11.
Z. Zhou, M. M. Rahman Siddiquee, N. Tajbakhsh,
and J. Liang, “Unet++: A nested u-net architecture
for medical image segmentation,” in Deep Learning in
Medical Image Analysis and Multimodal Learning for
Clinical Decision Support, Springer, 2018, pp. 3–11.
DOI: https://doi.org/10.1007/978-3-030-00889-5 1.
H. Huang, L. Lin, R. Tong, et al., “Unet 3+: A full-
scale connected unet for medical image segmenta-
tion,” in ICASSP 2020-2020 IEEE international con-
ference on acoustics, speech and signal processing
(ICASSP), IEEE, 2020, pp. 1055–1059. DOI: 10.1109/
ICASSP40776.2020.9053405.
O. Oktay, J. Schlemper, L. L. Folgoc, et al., “Attention
u-net: Learning where to look for the pancreas,” arXiv
preprint arXiv:1804.03999, 2018. DOI: https://doi.org/
48550/arXiv.1804.03999.
L. Mou, Y. Zhao, L. Chen, et al., “Cs-net: Channel and
spatial attention network for curvilinear structure seg-
mentation,” in Medical Image Computing and Computer
Assisted Intervention–MICCAI 2019, Springer, 2019,
pp. 721–730. DOI: https : / / doi . org / 10 . 1007 / 978 - 3 -
-32239-7 80.
J. Chen, Y. Lu, Q. Yu, et al., “Transunet: Transformers
make strong encoders for medical image segmentation,”
arXiv preprint arXiv:2102.04306, 2021. DOI: https : / /
doi.org/10.48550/arXiv.2102.04306.
A. Hatamizadeh, Y. Tang, V. Nath, et al., “Unetr:
Transformers for 3d medical image segmentation,” in
Proceedings of the IEEE/CVF winter conference on
applications of computer vision, 2022, pp. 574–584.
DOI: 10.1109/WACV51458.2022.00181.