A Memetic Genetic Particle Swarm Optimization for Druglike Molecule Discovery
Keywords:
De-Novo Drug Discovery, Druglike Molecules Design, Memetic Approach, MetaheuristicsAbstract
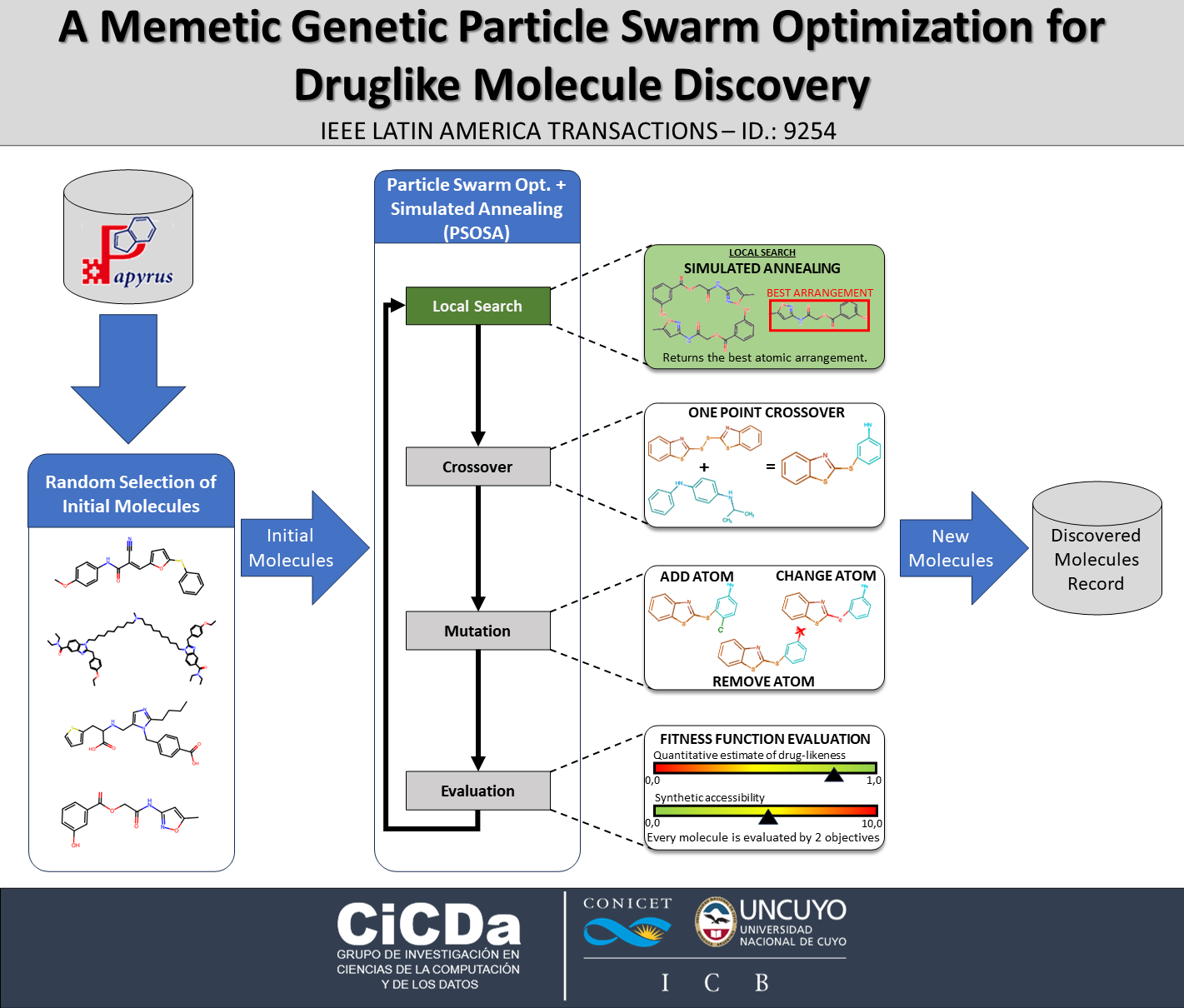
Given the vast and complex chemical search space, developing new techniques for identifying promising ligands that satisfy multiple objectives is highly desirable to reduce the costs and times required for effective drug discovery. Neural networks are frequently employed for this task, but they tend to generate molecules that are invalid both chemically and syntactically. As an alternative, metaheuristics have emerged as promising approaches, delivering notable results with reasonable computational costs. However, they often suffer from information loss during the process, leading to poor quality generations. In this work, we introduce a novel memetic algorithm that hybridizes Particle Swarm Optimization with Simulated Annealing. This approach aims to improve the balance between exploration and exploitation in the de-novo drug discovery process, ensuring that promising molecules are not overlooked during generation steps. We compare our approach against six state-of-the-art algorithms, and the results demonstrate that our algorithm enhances molecule generation quality, showing an increased diversity and improved chemical properties of the resulting ligands.
Downloads
References
C. M. Dobson, “Chemical space and biology,” Nature, vol. 432, no.
, pp. 824–828, Dec. 2004. doi: 10.1038/nature03192
T. Pereira, M. Abbasi et al., “Optimizing blood–brain barrier permeation through deep reinforcement learning for de novo drug design,”
Bioinformatics, vol. 37, no. Supplement_1, pp. i84–i92, Jul. 2021. doi:
1093/bioinformatics/btab301
X. Liu, K. Ye et al., “DrugEx v2: de novo design of drug molecules by
Pareto-based multi-objective reinforcement learning in polypharmacology,” J. Cheminf., vol. 13, no. 1, p. 85, Dec. 2021. doi: 10.1186/s13321-021-00561-9
A. Zhavoronkov, Y. A. Ivanenkov et al., “Deep learning enables rapid
identification of potent DDR1 kinase inhibitors,” Nat. Biotechnol.,
vol. 37, no. 9, pp. 1038–1040, Sep. 2019. doi: 10.1038/s41587-019-0224-x
B. Sanchez-Lengeling, C. Outeiral et al., “Optimizing distributions
over molecular space. An Objective-Reinforced Generative Adversarial
Network for Inverse-design Chemistry (ORGANIC),” Aug. 2017.
F. Zhu, X. X. Li et al., “Clinical Success of Drug Targets Prospectively
Predicted by In Silico Study,” Trends in Pharmacological Sciences,
vol. 39, no. 3, pp. 229–231, Mar. 2018. doi: 10.1016/j.tips.2017.12.002
D. Weininger, “SMILES, a chemical language and information system.
Introduction to methodology and encoding rules,” J. Chem. Inf.
Comput., vol. 28, no. 1, pp. 31–36, Feb. 1988. doi: 10.1021/ci00057a005
L. Schoenmaker, O. J. M. Béquignon et al., “UnCorrupt SMILES: a
novel approach to de novo design,” J. Cheminf., vol. 15, p. 22, Feb.
doi: 10.1186/s13321-023-00696-x
Y. Kwon and J. Lee, “MolFinder: an evolutionary algorithm for the
global optimization of molecular properties and the extensive exploration
of chemical space using SMILES,” J. Cheminf., vol. 13, no. 1, p. 24,
Mar. 2021. doi: 10.1186/s13321-021-00501-7
R. Devi, S. S. Sathya, and M. Coumar, “Evolutionary algorithms for
de novo drug design – A survey,” Applied Soft Computing, vol. 27, pp.
–552, Feb. 2015. doi: 10.1016/j.asoc.2014.09.042
S. Luukkonen, H. W. van den Maagdenberg et al., “Artificial intelligence
in multi-objective drug design,” Curr. Opin. Struct. Biol., vol. 79, p.
, Apr. 2023. doi: 10.1016/j.sbi.2023.102537
N. Jain, A. Hornback, and M. D. Wang, “EMxDesign: A Genetic
Algorithm for High Affinity Drug Design,” in Proceedings of the Genetic
and Evolutionary Computation Conference Companion, ser. GECCO ’24
Companion. New York, NY, USA: Association for Computing Machinery, Aug. 2024. doi: 10.1145/3638530.3654423. ISBN 9798400704956
pp. 439–442.
T. M. Shami, A. A. El-Saleh et al., “Particle Swarm Optimization: A
Comprehensive Survey,” IEEE Access, vol. 10, pp. 10 031–10 061, 2022.
doi: 10.1109/ACCESS.2022.3142859
M. Hartenfeller, E. Proschak et al., “Concept of Combinatorial De Novo
Design of Drug-like Molecules by Particle Swarm Optimization,” Chem.
Biol. Drug. Des., vol. 72, no. 1, pp. 16–26, 2008. doi: 10.1111/j.1747-0285.2008.00672.x
R. Winter, F. Montanari et al., “Efficient multi-objective molecular
optimization in a continuous latent space,” Chem. Sci., vol. 10, no. 34,
pp. 8016–8024, Aug. 2019. doi: 10.1039/C9SC01928F
C. Cotta, L. Mathieson, and P. Moscato, Memetic Algorithms. Cham:
Springer International Publishing, 2017, pp. 1–32. ISBN 978-3-319-07153-4
P. Moscato and C. Cotta, “An Accelerated Introduction to Memetic
Algorithms,” in Handbook of Metaheuristics, M. Gendreau and J.-Y.
Potvin, Eds. Cham: Springer International Publishing, 2019, vol. 272,
pp. 275–309. ISBN 978-3-319-91085-7 978-3-319-91086-4
V. Bagal, R. Aggarwal et al., “MolGPT: Molecular Generation Using a
Transformer-Decoder Model,” J. Chem. Inf. Model, vol. 62, no. 9, pp.
–2076, May 2022. doi: 10.1021/acs.jcim.1c00600
“CDDLeiden/DrugEx,” May 2024. [Online]. Available: https://github.com/CDDLeiden/DrugEx
A. Benítez-Hidalgo, A. J. Nebro et al., “jMetalPy: A Python framework
for multi-objective optimization with metaheuristics,” Swarm Evol. Comput., vol. 51, p. 100598, Dec. 2019. doi: 10.1016/j.swevo.2019.100598
O. J. M. Béquignon, B. J. Bongers et al., “Papyrus: a large-scale curated
dataset aimed at bioactivity predictions,” J. Cheminf., vol. 15, no. 1, p. 3,
Jan. 2023. doi: 10.1186/s13321-022-00672-x
A. P. Bento, A. Hersey et al., “An open source chemical structure
curation pipeline using RDKit,” J. Cheminf., vol. 12, no. 1, p. 51, Sep.
doi: 10.1186/s13321-020-00456-1
V. D. Mouchlis, A. Afantitis et al., “Advances in De Novo Drug Design:
From Conventional to Machine Learning Methods,” Int. J. Mol. Sci.,
vol. 22, no. 4, p. 1676, Jan. 2021. doi: 10.3390/ijms22041676
C. A. Lipinski, F. Lombardo et al., “Experimental and computational
approaches to estimate solubility and permeability in drug discovery
and development settings,” Adv. Drug Deliv. Rev., vol. 23, no. 1, pp.
–25, Jan. 1997. doi: 10.1016/S0169-409X(96)00423-1